|
CRISPR/Cas9 targeting events cause complex deletions and insertions at 17 sites in the mouse genome. CRISPR off-target detection with DISCOVER-seq. The next generation of CRISPR-Cas technologies and applications. Technologies and computational analysis strategies for CRISPR applications. Cornerstones of CRISPR-Cas in drug discovery and therapy. CRISPR-based technologies for the manipulation of eukaryotic genomes. 
Biology and applications of CRISPR systems: harnessing nature’s toolbox for genome engineering. Development and applications of CRISPR-Cas9 for genome engineering. The entire protocol from genomic DNA extraction to OnTE exclusion can be performed in 6–9 d. All steps are optimized to maximize simplicity and minimize cost to promote wide dissemination and applicability across the field. Second, occurrence of LOH around the edited locus is detected by genotyping neighbouring single-nucleotide polymorphisms (SNPs), using either a Sanger sequencing-based method or SNP microarrays. This combination of genotyping and quantitation makes it possible to exclude clones with monoallelic OnTEs and hemizygous editing, which are often mischaracterized as correctly edited in standard Sanger sequencing. This is achieved by subjecting genomic DNA first to quantitative genotyping PCR (qgPCR), which determines the number of intact alleles at the target site using the same PCR amplicon that has been optimized for genotyping. 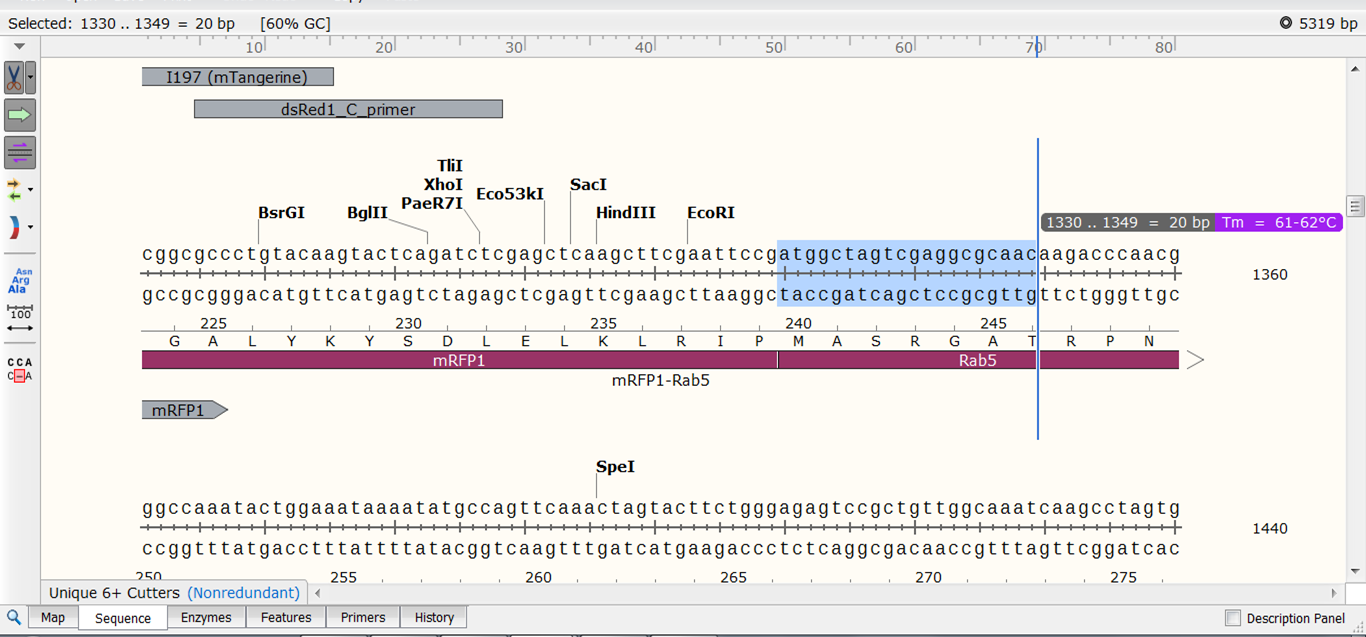
Our protocol enables identification of OnTEs such as large deletions, large insertions, rearrangements or loss of heterozygosity (LOH). Here we describe a broadly applicable framework for detecting OnTEs in genome-edited cells and organisms after non-homologous end joining-mediated and homology-directed repair-mediated editing. By altering gene expression of targeted or neighbouring genes, OnTEs can severely affect phenotypes of CRISPR-edited cells and organisms and thus lead to data misinterpretation, which can undermine the reliability of CRISPR-based studies. However, recent reports indicate widespread occurrence of unintended CRISPR-induced on-target effects (OnTEs) at the edited site in mice and human induced pluripotent stem cells (iPSCs) that escape standard quality controls. The recent CRISPR revolution has provided researchers with powerful tools to perform genome editing in a variety of organisms. Override the reading frame saved with the. Override the minimum ORF length saved with the. Override enzyme set to use saved with the. “ “.Īdds mouse handlers in the hittest region of the DOM. To clear the title completely from display, set the value to one space, e.g. If the subtitle is not defined or set to an empty string, then the default title (the number of base pairs) is used. “ “.ĭefines the subtitle to use for the map. If the title is not defined or set to an empty string, then the default title (the filename without the extenion) is used. 
Useful when supporting multiple maps or sequences on the same page.ĭefines the title to use for the map. User defined attribute used to identify a particular map or sequence view. Useful when supporting multiple maps or sequences on the same page Removes extra SVG attributes that are not necessary for correctly rendering the SVG. Instead use the JS file in /opt/gslbiotech/snapgene-server/examples/js/map-seq-interactivity.jsįile to text fragment of the javascript code required for hover/selection (dynamic part).įile to text fragment of the css code required for proper rendering of the SVG DOM. NOTE: this parameter is deprecated - there is no need to generate JS files using generateSVGMap. The actual size might be different because the map generation code performs extra logic to ensure the text graphical elements are all visibleįile to text fragment of the svg objects and tooltip DOM objectsįile to text fragment of the javascript code required for hover/selection (static part) the static part can be shared for all maps for the same version of snapgene server. Set the requested size of the map in width x height. 
True for linear map, false for circular map
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
